54. A Dual-Model Machine Learning Framework for Interpretable Design and Ensemble Prediction of C-Amidated Antimicrobial Peptides
Abstract:
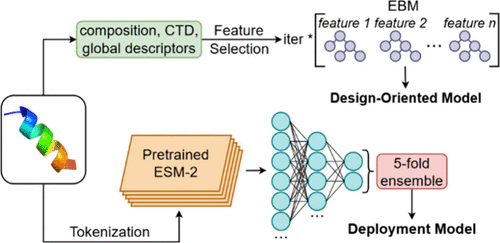
Antimicrobial peptides (AMPs) offer promising alternatives to conventional antibiotics, yet most predictive models fail to account for chemical modifications that influence real-world efficacy. Among these, C-terminal amidation is a widely adopted and effective strategy that improves structural stability, membrane interaction, and protease resistance. In this study, we established an integrated framework for the design and prediction of C-terminal amidated AMPs targeting Escherichia coli. Our approach combined a design-oriented model based on an interpretable Explainable Boosting Machine (EBM), which extracts actionable sequence-level design rules, together with a reliable deployment model, built on a fine-tuned ESM2 deep learning architecture. The resulting tool, CAmidPred, enables both predictive classification and amino acid pattern analysis with outputs examined in relation to published alanine-scanning experiments. Using these models, we identified a pardaxin variant with improved activity against E. coli, demonstrating the practical utility of the dual-model framework in targeted AMP design.
Le, D.; Zhu, Y.; Zhang, T.; Li, W.; Hung, A.; Houshyar, S.; Le, T., (2026) A Dual-Model Machine Learning Framework for Interpretable Design and Ensemble Prediction of C-Amidated Antimicrobial Peptides, ACS Appl. Mater. Interfaces, 18, 10, 14739–14754, DOI: 10.1021/acsami.6c00110.